Abstract
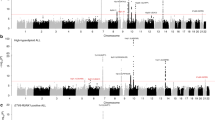
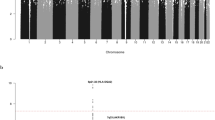
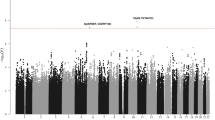
Recent genome-wide association studies (GWAS) have provided the first unambiguous evidence that common genetic variation influences the risk of childhood B-cell precursor acute lymphoblastic leukemia (BCP-ALL), identifying risk single-nucleotide polymorphisms (SNPs) localizing to 7p12.2, 9p21.3, 10q21.2 and 14q11.2. The testing of SNPs individually for an association in GWA studies necessitates the imposition of a very stringent P-value to address the issue of multiple testing. While this reduces false positives, real associations may be missed and therefore any estimate of the total heritability will be negatively biased. Using GWAS data on 823 BCP-ALL cases by considering all typed SNPs simultaneously, we have calculated that 24% of the total variation in BCP-ALL risk is accounted for common genetic variation (95% confidence interval 6–42%). Our findings provide support for a polygenic basis for susceptibility to BCP-ALL and have wider implications for future searches for novel disease-causing risk variants.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Greaves M . Infection, immune responses and the aetiology of childhood leukaemia. Nat Rev Cancer 2006; 6: 193–203.
Kaatsch P . Epidemiology of childhood cancer. Cancer Treat Rev 2010; 36: 277–285.
Hodgson S, Maher E . A Practical Guide to Human Cancer Genetics. Cambridge University Press: Cambridge, 1999, pp 116–122.
Papaemmanuil E, Hosking FJ, Vijayakrishnan J, Price A, Olver B, Sheridan E et al. Loci on 7p12.2, 10q21.2 and 14q11.2 are associated with risk of childhood acute lymphoblastic leukemia. Nat Genet 2009; 41: 1006–1010.
Trevino LR, Yang W, French D, Hunger SP, Carroll WL, Devidas M et al. Germline genomic variants associated with childhood acute lymphoblastic leukemia. Nat Genet 2009; 41: 1001–1005.
Sherborne AL, Hosking FJ, Prasad RB, Kumar R, Koehler R, Vijayakrishnan J et al. Variation in CDKN2A at 9p21.3 influences childhood acute lymphoblastic leukemia risk. Nat Genet 2010; 42: 492–494.
Fletcher O, Houlston RS . Architecture of inherited susceptibility to common cancer. Nat Rev Cancer 2010; 10: 353–361.
Lee SH, Wray NR, Goddard ME, Visscher PM . Estimating missing heritability for disease from genome-wide association studies. Am J Hum Genet 2011; 88: 294–305.
UK Childhood Cancer Study Investigators. The United Kingdom Childhood Cancer Study: objectives, materials and methods. Br J Cancer 2000; 82: 1073–1102.
Power C, Elliott J . Cohort profile: 1958 British birth cohort (National Child Development Study). Int J Epidemiol 2006; 35: 34–41.
Tomlinson I, Webb E, Carvajal-Carmona L, Broderick P, Kemp Z, Spain S et al. A genome-wide association scan of tag SNPs identifies a susceptibility variant for colorectal cancer at 8q24.21. Nat Genet 2007; 39: 984–988.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 2007; 81: 559–575.
Yang J, Benyamin B, McEvoy BP, Gordon S, Henders AK, Nyholt DR et al. Common SNPs explain a large proportion of the heritability for human height. Nat Genet 2010; 42: 565–569.
Yang J, Lee SH, Goddard ME, Visscher PM . GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet 2011; 88: 76–82.
Wray NR, Yang J, Goddard ME, Visscher PM . The genetic interpretation of area under the ROC curve in genomic profiling. PLoS Genet 2010; 6: e1000864.
Hemminki K, Jiang Y . Risks among siblings and twins for childhood acute lymphoid leukaemia: results from the Swedish Family-Cancer Database. Leukemia 2002; 16: 297–298.
Zuk O, Hechter E, Sunyaev SR, Lander ES . The mystery of missing heritability: Genetic interactions create phantom heritability. Proc Natl Acad Sci USA 2012; 109: 1193–1198.
Sherborne AL, Hemminki K, Kumar R, Bartram CR, Stanulla M, Schrappe M et al. Rationale for an international consortium to study inherited genetic susceptibility to childhood acute lymphoblastic leukemia. Haematologica 2011; 96: 1049–1054.
Spencer CC, Su Z, Donnelly P, Marchini J . Designing genome-wide association studies: sample size, power, imputation, and the choice of genotyping chip. PLoS Genet 2009; 5: e1000477.
Acknowledgements
Leukaemia and Lymphoma Research (UK) and the Kay Kendall Leukaemia Fund provided principal funding for the study. Support from Cancer Research UK (C1298/A8362 supported by the Bobby Moore Fund) is also acknowledged. This study made use of control genotyping data generated by the Wellcome Trust Case–Control Consortium. We acknowledge the use of genotype data from the British 1958 Birth Cohort DNA collection, funded by the Medical Research Council grant G0000934 and the Wellcome Trust grant 068545/Z/02. A full list of the investigators who contributed to the generation of the data is available from www.wtccc.org.uk. Funding for the project was provided by the Wellcome Trust under award 076113 and 085475. We are grateful to all the patients and individuals for their participation and we would also like to thank the clinicians, other hospital staff and study staff who contributed to the blood sample and data collection for this study.
WEB ADDRESSES
The URLs for data presented herein are as follows: PLINK: http://pngu.mgh.harvard.edu/purcell/plink/ British 1958 birth cohort: http://www.b58cgene.sgul.ac.uk Ilumina: http://www.illumina.com/ Genome-wide Complex Trait Analysis (GCTA): http://gump.qimr.edu.au/gcta/ Childhood Leukemia International Consortium: https://ccls.berkeley.edu/clic
AUTHOR CONTRIBUTIONS
RSH designed the study; RSH and VE-M drafted the manuscript; VE-M and FJH performed statistical analyses; EP oversaw laboratory analyses; ES and SEK performed curation and sample preparation of MRC ALL 97 trial samples; TL and ER managed and maintained UKCCS sample data; MT performed curation and sample preparation of UKCCS samples; JMA and JAEI performed ascertainment, curation and sample preparation of Northern Institute for Cancer Research case series; RSH and MG obtained funding and designed parent project.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Enciso-Mora, V., Hosking, F., Sheridan, E. et al. Common genetic variation contributes significantly to the risk of childhood B-cell precursor acute lymphoblastic leukemia. Leukemia 26, 2212–2215 (2012). https://doi.org/10.1038/leu.2012.89
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2012.89
Keywords
This article is cited by
-
Polymorphism in IKZF1 gene affects clinical outcome in diffuse large B-cell lymphoma
International Journal of Hematology (2017)
-
Heritability estimates on Hodgkin’s lymphoma: a genomic- versus population-based approach
European Journal of Human Genetics (2015)
-
Excess congenital non-synonymous variation in leukemia-associated genes in MLL− infant leukemia: a Children’s Oncology Group report
Leukemia (2014)
-
Current evidence for an inherited genetic basis of childhood acute lymphoblastic leukemia
International Journal of Hematology (2013)