Abstract
Modeling empirical distributions of repeated counts with parametric probability distributions is a frequent problem when studying species abundance. One must choose a family of distributions which is flexible enough to take into account very diverse patterns and possess parameters with clear biological/ecological interpretations. The negative binomial distribution fulfills these criteria and was selected for modeling counts of marine fish and invertebrates. This distribution depends on a vector \(\left( K,\mathfrak {P}\right) \) of parameters, and ranges from the Poisson distribution (when \(K\rightarrow +\infty \)) to Fisher’s log-series, when \(K\rightarrow 0\). Moreover, these parameters have biological/ecological interpretations which are detailed in the literature and in this study. We compared three estimators of K, \(\mathfrak {P}\) and the parameter \(\alpha \) of Fisher’s log-series, following the work of Rao CR (Statistical ecology. Pennsylvania State University Press, University Park, 1971) on a three-parameter unstandardized variant of the negative binomial distribution. We further investigated the coherence underlying parameter values resulting from the different estimators, using both real count data collected in the Mauritanian Exclusive Economic Zone (MEEZ) during the period 1987–2010 and realistic simulations of these data. In the case of the MEEZ, we first built homogeneous lists of counts (replicates), by gathering observations of each species with respect to “typical environments” obtained by clustering the sampled stations. The best estimation of \(\left( K,\mathfrak {P}\right) \) was generally obtained by penalized minimum Hellinger distance estimation. Interestingly, the parameters of most of the correctly sampled species seem compatible with the classical birth-and-dead model of population growth with immigration by Kendall (Biometrika 35:6–15, 1948).






Similar content being viewed by others
References
Aidoo EN, Mueller U, Goovarets P, Hyndes GA (2015) Evaluation of geostatistical estimators and their applicability to characterise the spatial patterns of recreational fishing catch rates. Fish Res 168:20–32
Anscombe FJ (1950) Sampling theory of the negative binomial and logarithmic series distributions. Biometrika 36:358–382
Basu A, Shioya H, Park C (2011) Statistical inference. The minimum distance approach, vol 120., Monographs on statistics and applied probabilityChapman & Hall/CRC, Boca Raton
Beran RJ (1977) Minimum Hellinger distance estimates for parametric models. Ann Stat 5:445–463
Bergerard P, Domain F, Richer de Forges B (1983) Evaluation par chalutage des ressources démersales du plateau continental mauritanien. Bull. Centr. Nat. Rech. Océanogr.et des Pêches, Nouadhibou 11:217–290
Bliss CI, Fisher RA (1953) Fitting the negative binomial distribution to biological data. Biometrics 9:176–200
Boswell MT, Patil GP (1970) Chance mechanisms generating the negative binomial distribution. In: Patil GP (ed) Random counts in models and structures, vol 1. Pensylvania State University Press, University Park, pp 3–22
Boswell MT, Patil GP (1971) Chance mechanisms generating logarithmic series distribution used in the analysis of number of species and individuals. In: Patil G, Pielou E, Waters W (eds) Statistical ecology, vol 1. Pennsylvania State University Press, University Park, pp 99–130
Chen J, Rubin H (1986) Bounds for the difference between median and mean of Gamma and Poisson distributions. Stat Probab Lett 4:281–283
Cohen JE (1972) Markov population processes as models of primate social and population dynamics. Theor Popul Biol 3(2):119–134
Dabo-Niang S, Ferraty F, Vieu P (2007) On the using of modal curves for radar waveforms classication. Comput Stat Data Anal 51(10):4878–4890
Dabo-Niang S, Yao AF, Pischedda L, Cuny P, Gilbert F (2010) Spatial mode estimation for functional random fields with application to bioturbation problem. Stoch Environ Res Risk Assess 24(4):487–497
Dabo-Niang S, Hamdad L, Ternynck C, Yao AF (2014) A kernel spatial density estimation allowing for the analysis of spatial clustering. Application to Monsoon Asia Drought Atlas data. Stoch Environ Res Risk Assess 28:2075–2099
Di Salvo F, Ruggieri M, Plaia A (2015) Functional principal components analysis for multivariate multidimensional environmental data. Environ Ecol Stat 22(4):739–757
Diserud OH (2001) Detecting changes in diversity in a fluctuating environment based on simulation of stochastic processes. Oceanol Acta 24(5):505–517
Domain F (1986) Evaluation par chalutage des ressources démersales du plateau continental mauritanien. In: Josse E et Garcia S (eds). Description et évaluation des ressources halieutiques de la ZEE mauritanienne. Rapport du Groupe de travail CNROP/FAO/ORSTOM, Nouadhibou, Mauritanie, 16–27 septembre 1985, COPACE/PACE Séries 86/37: 248–273
Elliot JM (1979) Some methods for the statistical analysis of samples of benthic invertebrates, 2nd edn. Freshwater Biological Association Scientific Publication No. 25, Ambleside
Ferraty F, Vieu P (2006) Nonparametric functional data analysis. Theory and practice, Springer series in statistics. Springer, New York
Fisher RA, Corbet A, Williams CB (1943) The relation between the number of species and the number of individuals in a random sample of an animal population. J Anim Ecol 12:42–58
Gaertner JC, Mazouni N, Sabatier R, Millet B (1999) Spatial structure and habitat associations of demersal assemblages in the Gulf of Lions: a multicompartimental approach. Mar Biol 135:199–208
Gasser T, Hall P, Presnell B (1998) Nonparametric estimation of the mode of a distribution of random curves. J R Stat Soc B 60(4):681–691
Gibbs AL, Su FE (2002) On choosing and bounding probability metrics. Int Stat Rev 70(3):419–435
Hooten MB, Wikle C (2008) A hierarchical Bayesian non-linear spatio-temporal model for the spread of invasive species with application to the Eurasian Collared-Dove. Environ Ecol Stat 15:59–70
Johnson AF, Jenkins SR, Hiddink JG, Hinz H (2013) Linking temperate demersal fish species to habitat: scales, patterns and future directions. Fish Fish 14:256–280
Kendall DG (1948) On some modes of population growth leading to R. A. Fisher’s logarithmic series distribution. Biometrika 35:6–15
Kunin WE, Gaston KJ (1997) The biology of rarity. Causes and consequence of rare-common differences. Chapman & Hall, London
Lamouroux N, Capra H, Pouilly M, Souchon Y (1999) Fish habital preferences in large streams of southern France. Freshw Biol 42:673–687
Lewin W-C, Freyhof J, Huckstorf V, Mehner T, Wolter C (2010) When no catches matter: coping with zeros in environmental assessments. Ecol Indic 10:572–583
Magurran AE (2005) Species abundance distributions: pattern or processes? Funct Ecol 19:177–181
Mandal A, Basu A (2013) Minimum disparity estimation: improved efficiency through inliers modification. Comput Stat Data Anal 64:71–86
Manté C, Claudet J, Rebzani-Zahaf C (2003) Fairly processing rare and common species in multivariate analysis of ecological series. Application to macrobenthic communities from Algiers harbour. Acta Biotheor 51:277–294
Manté C, Durbec JP, Dauvin JC (2005) A functional data-analytic approach to the classification of species according to their spatial dispersion. Application to a marine macrobenthic community from the Bay of Morlaix (Western English Channel). J Appl Stat 32(8):831–840
Nielsen JR, Kristensen K, Lewy P, Bastardie F (2014) a statistical model for estimation of fish density including correlation in size, space, time and between species from research survey data. PloS One 9(6):1–15
O’Neil MF, Faddy MJ (2003) Use of binary and truncated negative binomial modelling in the analysis of recreational catch data. Fish Res 60:471–477
Quenouille MH (1949) A relation between the logarithmic. Poisson and negative binomial series. Biometrics 5(2):162–164
Rao CR (1962) Efficient estimates and optimum inference procedures in large samples (with discussion). J R Stat Soc, Ser B (Methodol) 24(1):46–72
Rao CR (1971) Some comments on the logarithmic series distribution in the analysis of insect trap data. In: Patil G, Pielou E, Waters W (eds) Statistical ecology, vol 1. Pennsylvania State University Press, University Park, pp 131–142
Serfling R (2004) Nonparametric multivariate descriptive measures based on spatial quantiles. J Stat Plan Inference 123:259–278
Simpson DG (1987) Minimum Hellinger distance estimation for the analysis of count data. J Am Stat Assoc 82(399):802–807
Taylor LR, Kempton RA, Woiwod IP (1976) Diversity statistics and the log-series model. J Anim Ecol 45(1):255–272
Taylor LR, Woiwod IP, Perry JN (1979) The negative binomial as a dynamic ecological model for aggregation, and the density dependence of K. J Anim Ecol 48:289–304
Vaudor L, Lamouroux N, Olivier JM (2011) Comparing distribution models for small samples of overdispersed counts of freshwater fish. Acta Oecol 37(3):170–178
Wang Y (1996) Estimation problems for the two-parameters negative binomial distribution. Stat Probab Lett 26:113–114
Warton DI (2005) Many zeros does not mean zero inflation: comparing the goodness-of-fit of parametric models to multivariate abundance data. Environmetrics 16:275–289
Watterson GA (1974) Models for the logarithmic species abundance distributions. Theor Popul Biol 6:217–250
Williams CB (1944) Some applications of the logarithmic series and the index of diversity to ecological problems. J Ecol 32(1):1–44
Williams CB (1947) The logarithmic series and its application to biological problems. J Ecol 34(2):253–272
Williams CB (1952) Sequences of wet and of dry days considered in relation to the logarithmic series. Q J R Meteorol Soc 73(335):91–96
Wyshak G (1974) Algorithm AS 68: a program for estimating the parameters of the truncated negative binomial. J R Stat Soc, Ser C (Appl Stat) 23(1):87–91
Acknowledgments
We thank the Mauritanian Institute of Oceanographic Research and Fisheries (IMROP) and the Department of Cooperation and Cultural Action of the Embassy of France in Mauritania for their support for this study. We also thank all scientists who contributed to field surveys and data collection, Jean-Pierre Durbec for helpful discussions, and anonymous reviewers for their constructive comments. We are very grateful to Starrlight Augustine for greatly improving the English text.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling Editor: Pierre Dutilleul.
Appendices
Appendix 1: Determination of a threshold for the truncated Hellinger distance
While results about the asymptotic distribution of our estimators abound, nothing is known about the distribution of the goodness-of-fit index [see Formula (9)]
where \(\left( \hat{K},\mathfrak {\hat{P}}\right) \) is an estimate of \(\left( K,\mathfrak {P}\right) \). In order to determine from \(d_{H}^{T}\) the species which were correctly fitted, we performed a Monte Carlo study. It consisted in generating a sample of the statistics (10) for each one of the three estimators used, from a population of “negative binomial species” similar to the genuine population of the C4 habitat considered as a reference structure. This study is detailed hereunder.
1.1 The reference distribution of \(\left( K,\mathfrak {P}\right) \)
We plotted on Fig. 7 the minimum \(PHD_{h}\) estimates of the vector \(\left( K,\mathfrak {P}\right) \) associated with the species collected in the C4 habitat. About 35.6 % of the species were associated with very small values of the first parameter (\(\hat{K}\le e^{-10}\)); discarding these species, we could fit a bi-dimensional log-normal distribution of parameters \(\left( \mu _{B},\Sigma _{B}\right) \) to the remaining vectors of estimates, whose confidence ellipsoids are also represented on Fig. 7. Neither \(Log\left( \hat{K}\right) \) nor \(Log\left( \mathfrak {\hat{P}}\right) \) strictly obeyed a normal distribution, but this model was retained for the sake of simplicity, since the corresponding 95 % confidence region widely covers the data (see Fig. 7). As for the discarded species, we postulated that \(Log\left( \mathfrak {\hat{P}}\right) \) could be considered as obeying some Gaussian distribution, \(\mathcal {N}\left( \mu _{S},\sigma _{S}\right) \).
1.2 Generating a “population” consistent with the reference distribution
To build a sample of \(d_{H}^{T}\left( TNBD\left( \hat{K},\mathfrak {\hat{P}}\right) ,\,TNBD\left( K,\mathfrak {P}\right) \right) \) for counts having the same overall characteristics as the C4 data, we generated random counts of 300 “NB species”, whose random parameters obeyed the mixture distribution
where \(\left( \mu _{S},\sigma _{S}\right) =\left( -0.701062,2.68525\right) \) and \(\left( \mu _{B},\Sigma _{B}\right) \) were estimated from the C4 data. In practice, the parameters \(\left( k,p\right) \) of each species were first drawn according to \(\mathcal {M}\); then a sample of \(\beta =3000\) (or \(\beta =6000\) when \(k\le e^{-8}\)) counts obeying \(NBD\left( k,p\right) \) were drawn. The simulated data were then processed the same way as the MEEZ ones.
In black: 50 and 95 % confidence ellipsoids for the reference distribution \(\mathcal {N}\left( \mu _{B},\Sigma _{B}\right) \); in gray: same confidence ellipsoids for the distribution \(\mathcal {N}\left( \widehat{\mu _{B}},\widehat{\Sigma _{B}}\right) \) obtained from the mixture distribution \(\mathcal {M}\). Dots correspond to the parameters of the “NB species”, estimated by PHD \(_{h}\)
On Fig. 8, we superimposed to the parameters of the species (estimated by \(PHD_{h}\)), confidence ellipsoids of the reference distribution \(\mathcal {LN}\left( \mu _{B},\Sigma _{B}\right) \) and of the distribution \(\mathcal {LN}\left( \widehat{\mu _{B}},\widehat{\Sigma _{B}}\right) \), whose parameters were estimated from the independent draws of \(\mathcal {M}\). It is worth noting that in this case, there was no significant difference between the empirical distribution of \(\left( \hat{K},\mathfrak {\hat{P}}\right) \) and the reference distribution \(\mathcal {LN}\left( \mu _{B},\Sigma _{B}\right) \) (P values: Cramer–Von Mises =0.544087; Pearson \(\chi ^{2}\) = 0.523489).
1.3 Results
About 30 % (93) of the “species” were observed \({<}6\) times, and discarded. The goodness-of-fit density estimates for the remaining ones are plotted on Fig. 9. The three estimators perform equally well in the case of genuine TNB distributions, while the goodness-of-fit by ULSD is very different. Among the 207 remaining species, only 3 % (6) were better fitted by ULSD, while the best estimator for TNBD was \(PHD_{h}\) (185 species: 89 % of the total), followed by MLE (10 species) and pseudo-MLE (6 species). Thus almost all “aggregative species” were discarded due to their rarity.
The quantile of order 0.95 of the goodness-of-fit associated with \(PHD_{h}\) was 0.531096; consequently, we considered that \(d_{H}^{T} = 0.53\) is an approriate threshold value that would not be exceeded by genuine negative binomial species. This threshold was used in Sect. 6.
Appendix 2: The Williams–Rao’s condition and the estimation of \(\beta \)
Notice that the equality \(\alpha =K\,\beta \) is implicit in Eq. (4), but that it not a constraint in the root finding of the pseudo-MLE system (6). Consequently, this relationship must be considered as a sign of consistency of the estimation of \(\left( K,\mathfrak {P,\alpha }\right) \). If in addition, the first condition of (5) is fulfilled, we should also have: \(\dfrac{\alpha _{LS}}{K\,\beta }\approx \dfrac{\alpha _{Rao}}{K\,\beta }\approx 1\). We investigated the validity of this relationship by performing 50 Monte Carlo experiments with “negative binomial species” random draws, in each one of four typical cases:
-
the “mean” one: \(\left( K,\mathfrak {P}\right) =\left( 1.193,73.15\right) \) is the mean of the bivariate Log-normal distribution fitting the C4 habitat non-aggregative species (see Sect. 5.2 and Appendix 1)
-
the “common” one: \(\left( K,\mathfrak {P}\right) =\left( 0.7767,14.43\right) \) is the spatial median (Serfling 2004) of the parameters of the simulated “NB species”; in this case, we chose \(\beta =10^{5}\) as the sample size of each one of the 50 simulations
-
an “aggregative” case: \(\left( K,\mathfrak {P}\right) =\left( 0.0001,14.43\right) \), with \(\beta =10^{7}\)
-
a “bell-shaped” case: \(\left( K,\mathfrak {P}\right) =\left( 10,14.43\right) \), with \(\beta =10^{4}\).
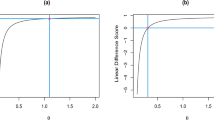
In these four cases, the best fit was obtained with \(PHD_{h}\), and we observed that \(\alpha _{LS}\) and \(\alpha _{Rao}\) could be considered as normally distributed (according to the Cramer-von Mises test), and that the mean of \(\alpha _{Rao}\) was always close to \(K\,\beta \) (T test), while the relationship \(\dfrac{\alpha _{LS}}{K\,\beta }\approx 1\) was verified only in the aggregative case (see Fig. 12). The equality \(\alpha _{LS}=K\,\beta \) was unacceptable in the “common” case (see Fig. 11), as well as in the bell-shaped one (Fig. 10) and in the “mean” case (not shown).
From another side, dealing with real data, we often have to face an additional estimation task: because of the dubious nature of the collected zeros (if there are), the true value of \(\beta \) is unknown and we only know the number \(\beta ^{+}\) of strictly positive counts. Then, it is classical to estimate \(\beta \) by:
where \(\left( \hat{K},\mathfrak {\hat{P}}\right) \) is an estimation of the TNBD parameters, obtained either from pseudo-MLE or \(PHD_{h}\). The bell-shaped case is problem-free, because the probability of zero is extremely weak.
Let us now examine the most interesting case: the “common” one. On Fig. 13, we plotted on the left panel KDE estimates of the 50 values of the expression (11) obtained with \(\left( \hat{K},\mathfrak {\hat{P},\hat{\alpha }}\right) =\left( K_{Rao},\mathfrak {P}_{Rao},\alpha _{Rao}\right) \) and divided by \(\beta \): they were very close to 1. On the right panel, we plotted the values of (11) obtained with \(\left( \hat{K},\mathfrak {\hat{P}}\right) =\left( K_{PHD_{h}},\mathfrak {P}_{PHD_{h}}\right) \). Thus, this figure shows that all estimates of \(\beta \) are excellent in the “common” case; as a consequence, the results above concerning the Williams–Rao’s condition (see Fig. 11) stay valid in this case.
Quite similar results were obtained in the “mean” case, as the reader can see on Fig. 14.
Things are very different in the aggregative situation, as the reader can see on Fig. 15: the estimator (11) based on \(\left( K_{Rao},\mathfrak {P}_{Rao},\alpha _{Rao}\right) \) strongly underestimates or overestimates \(\beta \). More precisely, in about 40 % of the samples, \(\beta \) was highly overestimated, which is natural, since \(\lim _{\hat{K}\rightarrow 0}\beta ^{\left( \hat{K},\mathfrak {\hat{P}}\right) }=\dfrac{\beta ^{+}}{\hat{K}\,\ln \left( 1+\mathfrak {\hat{P}}\right) }\). As for the estimator (11) associated with \(\left( \hat{K},\mathfrak {\hat{P}}\right) =\left( K_{PHD_{h}},\mathfrak {P}_{PHD_{h}}\right) \), it always underestimated \(\beta \). Consequently, when K is very small, the consistency of the estimations of the parameters of \(UNBD\left( K,\mathfrak {P,\alpha }\right) \) is questionable and the log-series model is probably more sound—even if the fit is not quite as good as with \(UNBD\left( K,\mathfrak {P,\alpha }\right) \). This is undoubtedly the meaning of Fig. 12.
Rights and permissions
About this article
Cite this article
Manté, C., Kidé, S.O., Yao-Lafourcade, AF. et al. Fitting the truncated negative binomial distribution to count data. Environ Ecol Stat 23, 359–385 (2016). https://doi.org/10.1007/s10651-016-0343-1
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10651-016-0343-1