5 minute read.Data citations and the eLife story so far
When we set up the eLife journal in 2012, we knew datasets were an important component of research content and decided to give them prominence in a section entitled ‘Major datasets’ (see images below). Within this section, major previously published and generated datasets are listed. We also strongly encourage data citations in the reference list.
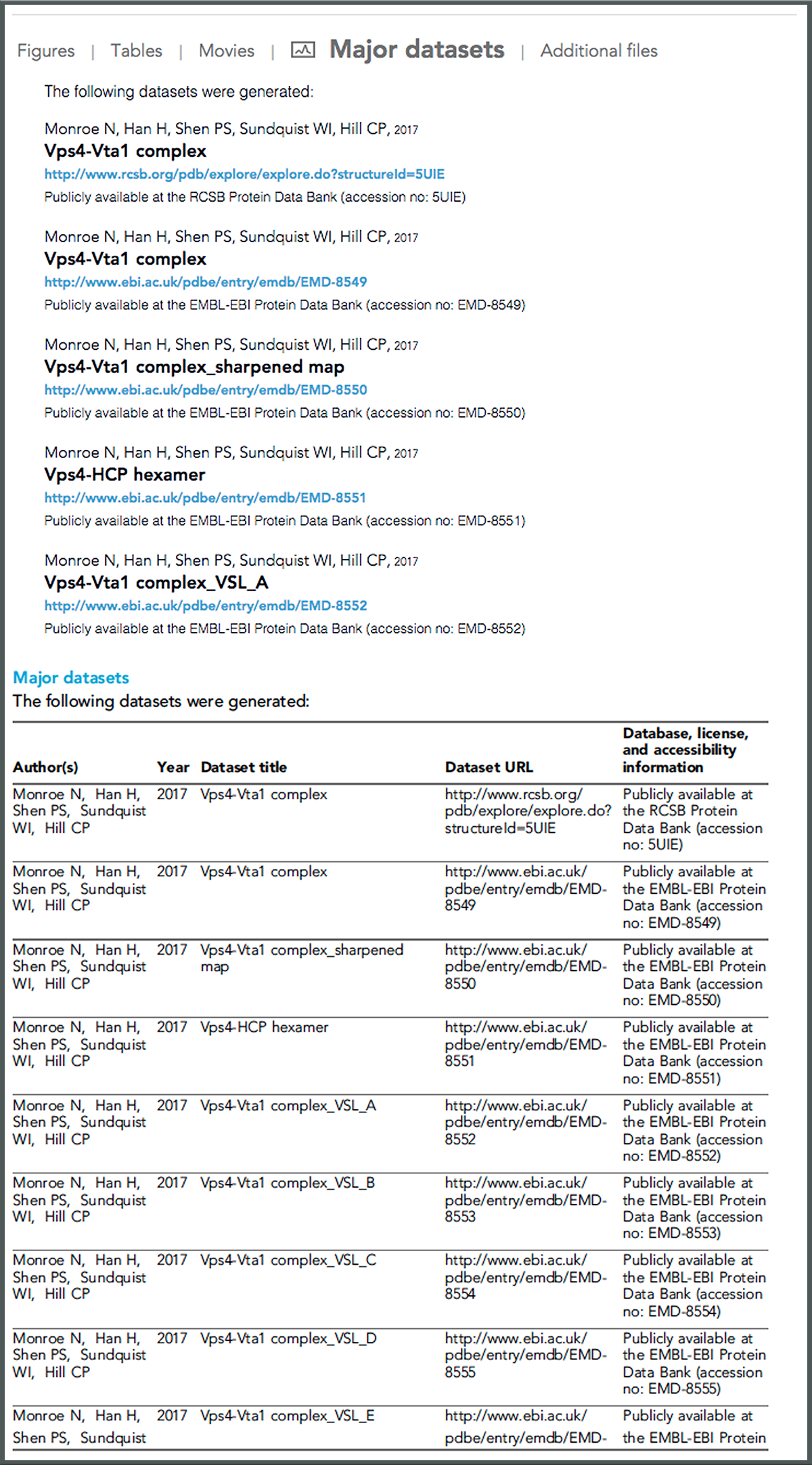
Major Datasets for “Structural basis of protein translocation by the Vps4-Vta1 AAA ATPase” by N. Monroe, H. Han, P. Shen, et. al.
Almost five years on and I feel we have still not cracked it! We have signed up to the Force11 data citation principles, which were published three years back; we have been actively involved in working groups of Force11 and others, for example the Data Citation Roadmap for Scientific Publishers and the JATS XML data citation recommendation of JATS4R. I am also currently working with other publishers to come up with recommended JATS XML tagging for data availability statements, which is easier said than done considering the nuances of dataset uses and also how different publishers approach this.
Added to this, there is still significant push-back from authors about putting all dataset citations in the reference list (for example, authors are concerned about self-citing by citing a dataset created as part of the research article; “dataset citations” that are in effect a link to a search results page on a database; and the necessitation of hundreds of reference entries if an author has used a large base for the research).
While eLife is very active in this space, and aims to arrange and mark up the datasets and citations produced by our authors in line with recommendations, the recommendations still have some gaps and the complete picture is not yet clear.
In late 2014, we brought in-house the process of depositing Crossref metadata (previously our online host did this for us). It gave us control of our processes and, at the time, we sent all the information we could to Crossref and have ensured our references are open and available in the Crossref public API. The code for this conversion process is all open-source and available for reuse. It can be found on GitHub. Since then, besides small improvements to the code and troubleshooting problems, we’ve not updated the code. I have been keeping a list of Crossref features and new deposit metadata we can add to our deposits, and now is the time for us to start working on this again.
One of the items we’ll be addressing is data citations.
The Crossref reference schema does not cater well for non-book or -journal content, and if an item does not have a DOI, the “reference” is not very useful because of the few tags available in the Crossref schema.
However, Crossref have introduced the relationship type to their schema, so data references can be well linked and mineable. As I see Crossref as a potential broker between publishers and data repositories in the future, using the relationship-type deposit for all datasets will assist this and also allow these data points to more easily be seen within the article Nexus framework (see the recent blog post, How do you deposit data citations?).
At eLife, we already distinguish between Dataset generated as part of research results (relationship type in the Crossref schema: “isSupplementedBy”) and Dataset produced by a different set of researchers or previously published (relationship type: “references”). Therefore, it will not be hard for us to convert all the information about data referencing that is within the dataset section into a relationship-type deposit in the conversion to Crossref XML.
We have also recently gone through an exercise of defining a set of rules for all our references and, of the 12 allowed types, one is data. The rules for Schematron (a rule-based validation language for making assertions about the presence or absence of patterns in XML trees; see also this useful article about Schematron on the JATS4R learning centre) have been written for the eLife ‘business’ rules. Subject to final testing, these will be integrated into our workflow (the Schematron is open source and available for reuse on GitHub, and we will also build an API for people to use the Schematron direct). This will allow us to easily identify all data references and convert them into relationship types in the XML delivered to Crossref. This way, they will not be lost in the references section of our deposits, but properly identified.
However, we do appreciate this will become harder for us as authors become more familiar with datasets as references, because we will not be able to identify the difference between generated and analysed datasets so easily.
The code developed and used to complete these conversions will, again, be on Github and open source, and we actively encourage the reuse of this.
While the industry is still working on the best way to deal with data and ensuring it is given the prominence it requires, we feel this is the best approach we can take. Nothing is forever and we can still change what we do in the future. The beauty of open-source code also means that if there is an alternative approach now or in the future, the code we wrote at eLife can be developed by someone else in the future and we can all benefit.
If you have any questions, please do not hesitate to contact us.







