Abstract
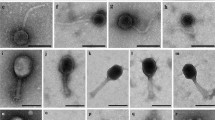
The complex cellulolytic microbial community of the horse intestines is a convenient model for studying the ecology of bacteriophages in natural habitats. Unlike the rumen of the ruminants, this community of the equine large intestine is not subjected to digestion. The inner conditions of the horse gut are much more stable in comparison to other mammals, due to the fact that the horse diet remains almost unchanged and the intervals between food consumption and defecation are much shorter than the whole digestive cycle. The results of preliminary analysis of the structure and dynamics of the viral community of horse feces, which combines direct and culture methods, are presented. In horse fecal samples, we detected more than 60 morphologically distinct phage types, the majority of which were present as a single phage particle. This indicates that the community includes no less than several hundreds of phage types. Some phage types dominated and constituted 5–11% of the total particle count each. The most numerous phage type had an unusual morphology: the tails of its members were extremely long (about 700 nm), flexible, and irretractable, while their heads were 100 nm in diameter. Several other phage types with similar but not identical properties were detected. The total coliphage plaque count of the samples taken from three animals revealed significant fluctuations in the phage titers. During the observation time, the maximum titer ranged within four orders of magnitude (103-107 plaque forming units (PFU)/g); the minimum titer ranged within two orders of magnitude. The samples contained two to five morphologically distinct and potentially competitive coliphage types, specific to a single Escherichia coli strain.
Similar content being viewed by others
References
Weinbauer, M., Ecology of Procaryotic Viruses, FEMS Microbiol. Rev., 2004, vol. 28, no. 2, pp. 127–181.
Proctor, L.M. and Fuhrman, J.A., Viral Mortality of Marine Bacteria and Cyanobacteria, Nature, 1990, vol. 343, no. 6253, pp. 60–62.
Proctor, L., Advances in the Study of Marine Viruses, Microsc. Res. Tech., 1997, vol. 37, no. 2, pp. 136–161.
Edwards, R.A. and Rohwer, F., Viral Metagenomics, Nat. Rev. Microbiol., 2005, vol. 3, no. 6, pp. 504–510.
Hintz, H.F. and Cymbaluk, N.F., Horse Nutrition, Annu. Rev. Nutr., 1994, vol. 14, pp. 243–267.
Cann, J.A., Fandrich, S.E., and Heaphy, S., Analysis of the Virus Population Present in Equine Faeces Indicates the Presence of Hundreds of Uncharacterized Virus Genomes, Virus Genes, 2005, vol. 30, no. 2, pp. 151–156.
Havelaar, A.H., Furuse, K., and Hogeboom, W.M., Bacteriophages and Indicator Bacteria in Human and Animal Faeces, J. Appl. Bacteriol., 1986, vol. 60, no. 3, pp. 255–262.
Calci, K.R., Burkhardt III, W., Watkins W.D., Scott R.R. Occurrence of Male-Specific Bacteriophage in Feral and Domestic Animals Wastes, Human Feces, and Human-Associated Wastewaters, Appl. Environ. Microbiol., 1998, vol. 64, no. 12, pp. 5027–5029.
Daly, K., Stewart, C.S., Flint, H.J., and Shirazi-Beechy, S.P., Bacterial Diversity within the Equine Large Intestine As Revealed by Molecular Analyses of Cloned 16S rRNA Genes, FEMS Microbiol. Ecol., 2001, vol. 38, pp. 141–151.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © E.E. Kulikov, A.S. Isaeva, A.S. Rotkina, A.A. Manykin, A.V. Letarov, 2007, published in Mikrobiologiya, 2007, Vol. 76, No. 2, pp. 271–278.
Rights and permissions
About this article
Cite this article
Kulikov, E.E., Isaeva, A.S., Rotkina, A.S. et al. Diversity and dynamics of bacteriophages in horse feces. Microbiology 76, 236–242 (2007). https://doi.org/10.1134/S0026261707020166
Received:
Issue Date:
DOI: https://doi.org/10.1134/S0026261707020166